Unifying biomedical knowledgein a modern multimodal graph

- 65sources
- 18ontologies
- 10entity types
- 190,531nodes
- 27relation types
- 21,813,816edges
- 110,276,843properties
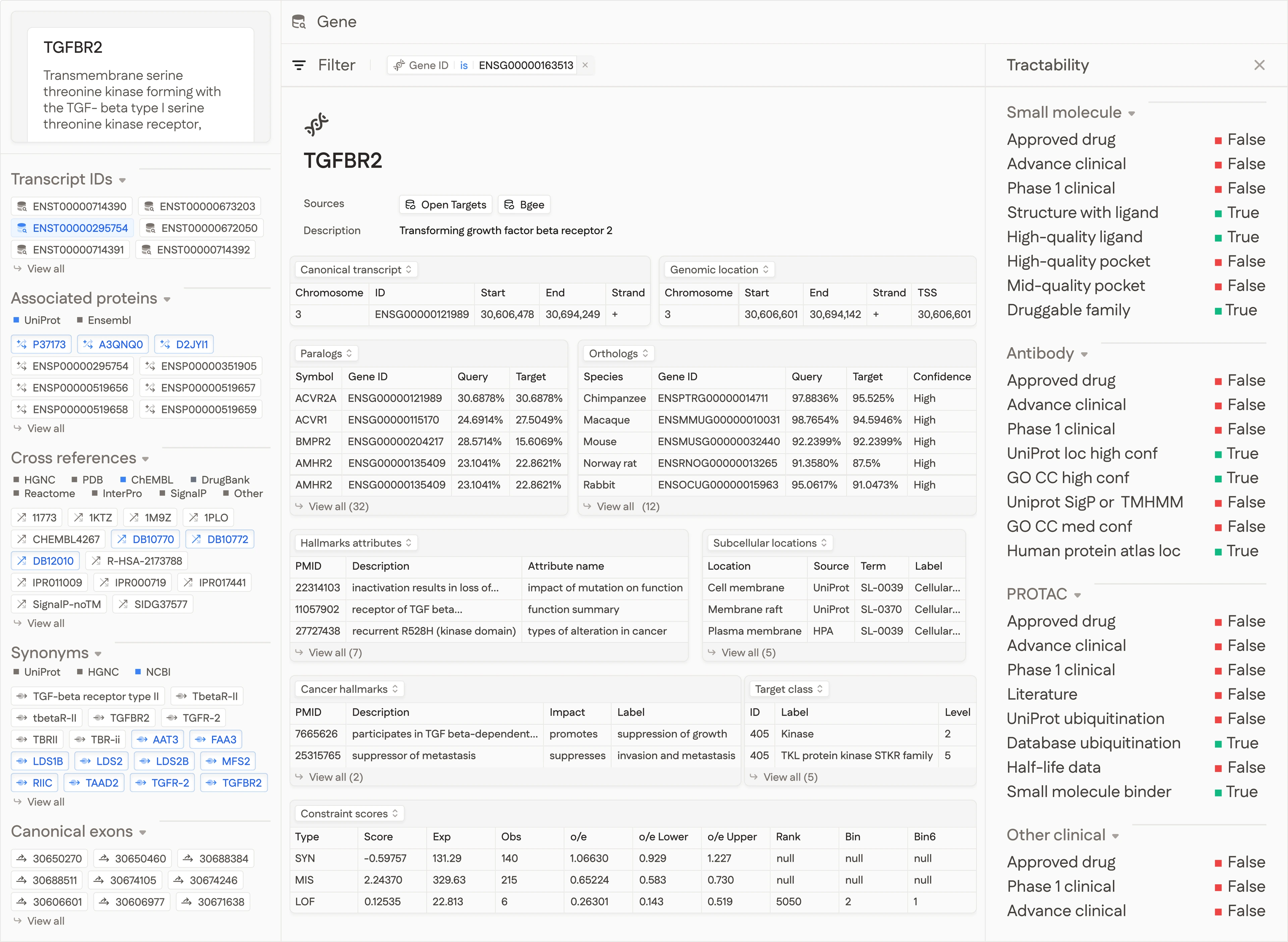
Rich strongly-typed properties
Every entity is enriched with structured properties for fine-grained analysis.

Graph Schema
idStringNode identifier in CURIE format (e.g. ENSG00000141510)
labelStringNode type abbreviation (GEN)
propertiesStructGene-specific properties
symbolStringOfficial HGNC gene symbol (e.g. TP53)
nameStringFull gene name
biotypeStringGene biotype (e.g. protein_coding, lncRNA)
genomic_locationStructChromosomal coordinates
chromosomeStringChromosome name
startInt64Start position (0-based)
endInt64End position
strandInt32Strand (+1 forward, -1 reverse)
transcription_start_siteInt64Transcription start site position
transcript_idsList[String]All associated Ensembl transcript IDs
function_descriptionsList[String]Functional descriptions
xrefsList[Struct]Cross-references to external databases
idStringExternal identifier
sourceStringDatabase name
sourcesStructProvenance of this node
directList[String]Datasets that directly contributed this entity
indirectList[String]Datasets that referenced this entity
Delightfully simple Python client
Install with one command and load the graph as Polars data frames or a NetworkX graph in a single line.
uv add optimuskg

Python Client
import optimuskg
# Download a specific file and store it locally
local_path = optimuskg.get_file("nodes/gene.parquet")
# Load a single Parquet file as a Polars DataFrame
drugs = optimuskg.load_parquet("nodes/drug.parquet")
# Load nodes and edges as Polars DataFrames
# Set lcc=True to load only the largest connected component
nodes, edges = optimuskg.load_graph(lcc=True)
# Load the graph as a NetworkX MultiDiGraph with metadata
# Set lcc=True to load only the largest connected component
G = optimuskg.load_networkx(lcc=True)
